Comparative In Silico Analysis of Respiratory Burst Oxidase Homologs (Rboh) Gene Family in Economically Significant Dicot Crop Plants
DOI:
https://doi.org/10.55627/pbiotech.003.02.1341Keywords:
Rboh gene family, Reactive oxygen species (ROS), Dicot crop plants, Genome-wide analysis, Plant stress resilienceAbstract
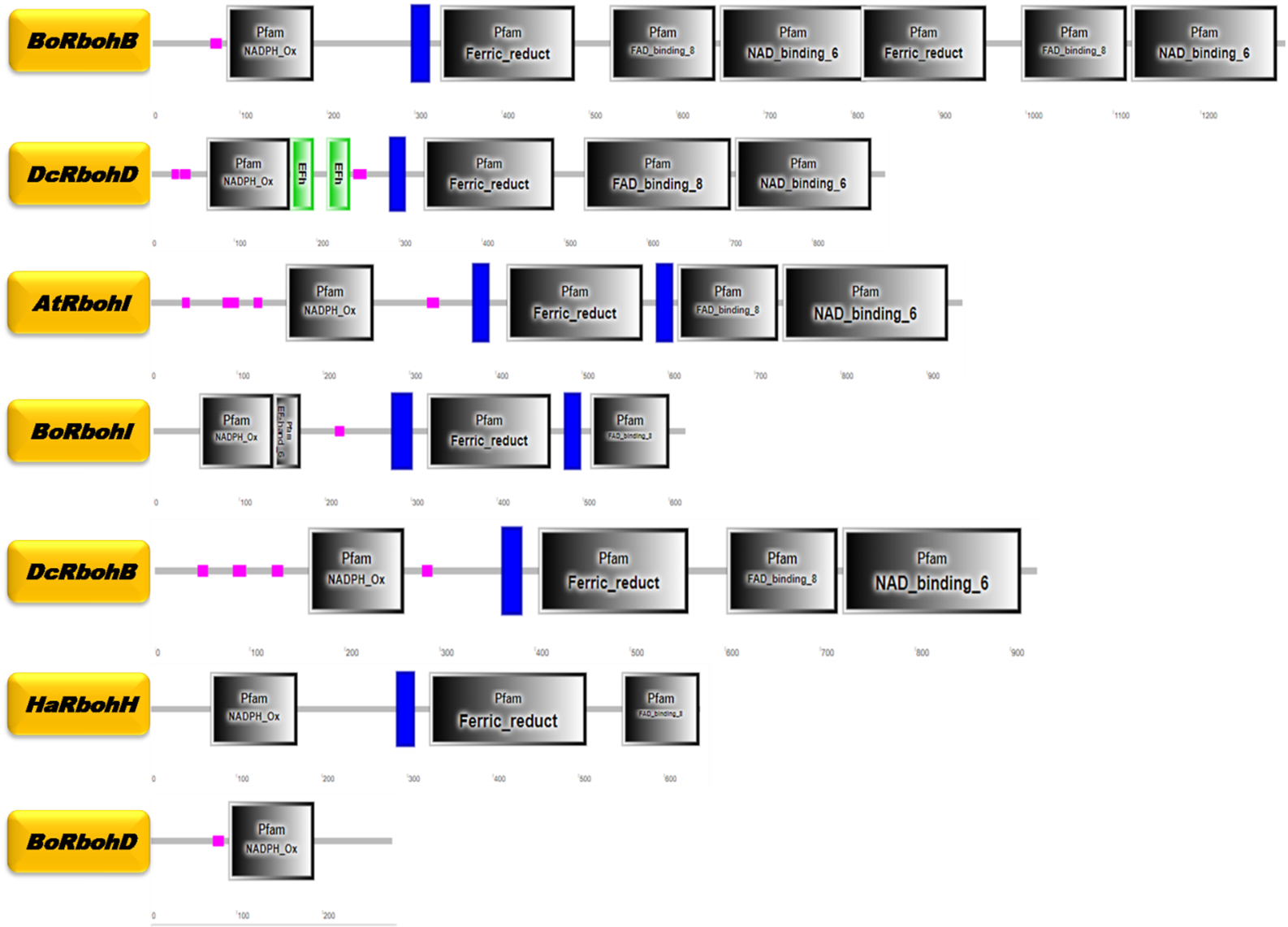
Reactive oxygen species (ROS), such as superoxide and hydrogen peroxide, play a pivotal role in the early defense mechanisms of plants against biotic and abiotic stresses. Among the key contributors to ROS production are respiratory burst oxidase homologs (Rbohs), plant-specific NADPH oxidases that regulate stress responses, signaling, development, and programmed cell death. While Rboh gene families have been extensively studied in model plants like Arabidopsis thaliana, their characterization in other dicot species remains limited. Here, we conducted a genome-wide analysis of Rboh genes in four agriculturally important dicot species: Brassica oleracea, Daucus carota, Helianthus annuus, and Capsicum annuum. Using Arabidopsis Rboh sequences as references, we identified 38 Rboh genes across these species, which were compared with the 10 known Arabidopsis Rboh genes. Chromosomal mapping, gene structure, conserved domain, and motif analyses revealed that Rboh genes are evolutionarily conserved, with gene numbers aligning closely to those in previously studied plants. Phylogenetic analysis grouped these genes into two major clades and six subgroups, reflecting shared evolutionary lineages and potential functional similarities. Variations in exon-intron structures and unique domain duplications or deletions indicated possible functional divergence among specific genes. Chromosomal mapping showed uneven distribution of Rboh genes within each genome, consistent with patterns observed in other plant species. This study enhances the current understanding of the Rboh gene family in dicots, providing a foundation for future research on their functional roles and potential applications in improving plant resilience through genetic engineering or breeding strategies.
References
Downloads
Published
Issue
Section
License
Copyright (c) 2025 Zeenat Niaz, Junaid Ul Hassan, Daraz Ahmad, Raheela Amin, Sobia Tariq, Abdul Haseeb, Adil Zahoor (Author)

This work is licensed under a Creative Commons Attribution 4.0 International License.





