Identification and Expression Profiling of the Dof Gene Family in Panicum virgatum Revealing Their Role in Abiotic Stress Tolerance
DOI:
https://doi.org/10.55627/pbiotech.003.03.1449Keywords:
Dof gene family, Genome-wide analysis, Panicum virgatum, Expression analysis, Abiotic stressesAbstract
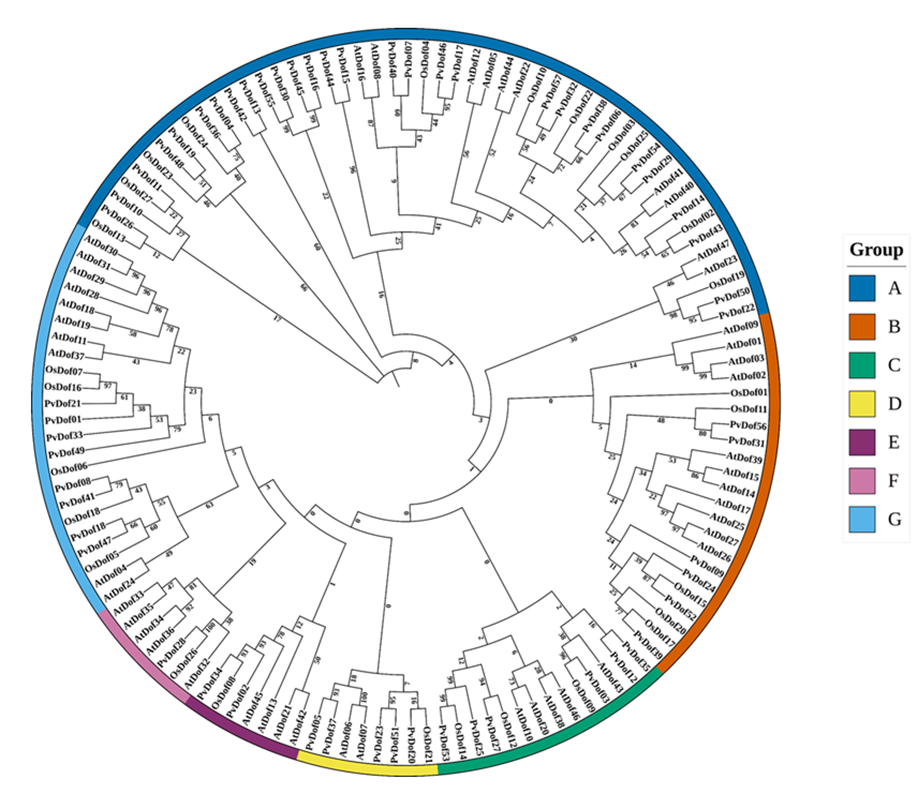
Switchgrass (Panicum virgatum) is a C4, perennial grass, with immense potential for biofuel production due to its adaptability, high biomass yield, and ecological benefits. Despite its agronomic importance, transcription factor regulation underlying stress responses in this crop remains unexplored. Therefore, we performed genome-wide analysis of Dof TF family in Panicum virgatum in which 57 non-redundant PvDof genes were identified and mapped across 17 chromosomes. Performing various bioinformatic analyses including phylogeny, motif, and gene structure, we classified these PvDofs into 7 phylogenetic groups (A-G) based on their conserved domains and sequence homology with Arabidopsis thaliana and Oryza sativa. Gene structure and conserved motif analyses demonstrated low intron counts and conserved functional motifs respectively, indicating functional conservation and potential involvement in stress signaling pathways. Promoter analysis uncovered 55 distinct cis-regulatory elements, including those associated with hormone signaling, light responsiveness and abiotic stress adaptation. Synteny analysis revealed more than 50 orthologous gene pairs between P. virgatum and A. thaliana, while 50 duplicated segments were identified in PvDof genes during collinearity analysis. Expression profiling under cadmium chloride exposure showed that PvDof06, PvDof32, PvDof38, and PvDof57 were strongly upregulated, whereas most other members were downregulated. Under phosphorus deficiency, PvDof32, PvDof38, and PvDof57 were induced in shoots, while PvDof05, PvDof06, and PvDof57 were highly expressed in roots, revealing tissue-specific regulation. These findings established a foundational framework for regulatory roles of Dof transcription factors in switchgrass and offer promising targets for genetic improvements of stress resilience in switchgrass through molecular breeding.
References
Downloads
Published
Issue
Section
License
Copyright (c) 2025 Mubashir Sharif, Muhammad Annas Shahid, Muzammal Majeed, Syeda Sahar Fatima, Saad Kamran, Rida Naseem, Mahad-Ur-Rehman, Ikhlas Shafique (Author)

This work is licensed under a Creative Commons Attribution 4.0 International License.





